Genotype Vs Phenotype: Examples And Definitions
Molly CampbellTechnology Networkseditorial policies
Complete the form below and we will email you a PDF version of“Genotype vs Phenotype: Examples and Definitions”
First Name*Would you like to receive further email communication from Technology Networks?Technology Networks Ltd. needs the contact information you provide to us to contact you about our products and services. You may unsubscribe from these communications at any time. For information on how to unsubscribe, as well as our privacy practices and commitment to protecting your privacy, check out our Privacy Policy
Any organism is a by-product of both its genetic makeup and the environment. To understand this in detail, we must first appreciate some basic genetic vocabulary and concepts. Here, we provide definitions for the terms genotype and phenotype, discuss their relationship and take a look at why and how we might choose to study them.
Identifying Genetic Variants Underlying Complex Traits
During 2007, the first wave of genome-wide association studies using tag SNPs resulted in the identification of common genetic variants associated with a broad range of common diseases and traits, including cancer, metabolic diseases, immune-mediated diseases and neurodegenerative diseases . The findings of these genome-wide scans can best be reviewed by discussing the results of studies investigating specific complex diseases and traits. Gout and its associated serum uric acid concentration has been studied in two genome-wide association studies , resulting in the identification of variants in the gene SLC2A9 . SLC2A9 variants were associated with high concentration of uric acid in the serum and the expression level of the isoform 2 of SLC2A9 was correlated with serum uric acid concentration . This isoform encodes the protein Glut9N, a putative fructose transporter expressed in kidney. As fructose is upstream in the pathway generating uric acid, an impaired expression of this protein possibly leads to the increased level of serum uric acid observed in gout .
Table 1 Genetic loci associated with disease and phenotypic variation
Evolutionary Origin Of Phenotype
The RNA world is the hypothesized pre-cellular stage in the evolutionary history of life on earth, in which self-replicating RNA molecules proliferated prior to the evolution of DNA and proteins. The folded three-dimensional physical structure of the first RNA molecule that possessed ribozyme activity promoting replication while avoiding destruction would have been the first phenotype, and the nucleotide sequence of the first self-replicating RNA molecule would have been the original genotype.
Recommended Reading: What Are Biological Indicators For Sterilization
Iculate Factorsmapping Genes Along Chromosomes
Mendelian research soon showed the independent assortment of factorsin Mendels experiments to be a special case, not a law.Departures from independent assortment of traits allowed theidentification of linkage groups, in which variants of two or moretraits co-occur, which eventually were shown to correspond to theproximity of their place or locus on distinct chromosomes.Johannsens resistance to the idea that discreteparticles of the chromosomes are bearers of specialparts of the whole inheritance was sharedby others , but such reservations did not hold backthe rise of Mendelianism to a dominant position in research intoheredity well before the material make-up and functioning of thoseparticlesthe geneswas revealed in the 1950s. Theparticulate view was affirmed by producing heritable alterations inphenotypes after bombarding organisms with high-energy ionizingradiation.
Difference Between Phenotype And Genotype
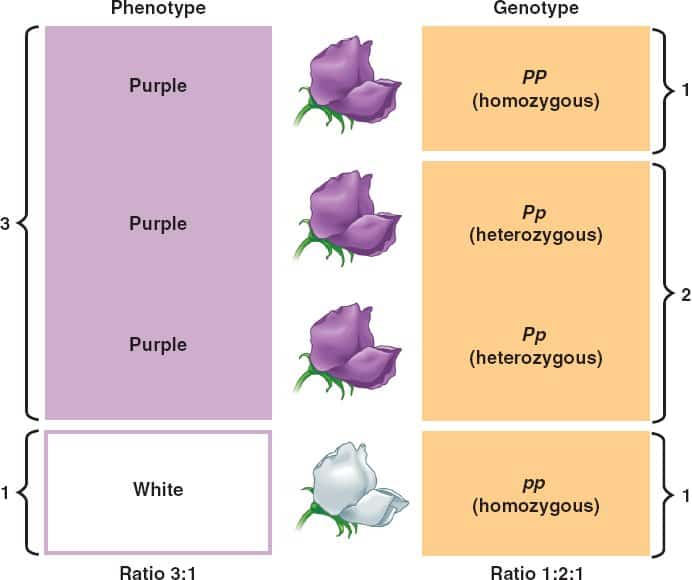
The genotype is a set of genes in the DNA which are responsible for the unique trait or characteristics. Whereas the phenotype is the physical appearance or characteristic of the organism. Thus, we can find the human genetic code with the help of their genotype. Organisms may look the same but still may not have the same genotype. One can determine the genotype by the biological tests. On the other hand, the phenotype is determined by an individuals genotype and can be expressed as genes or by the visible traits. Such traits are hair color or type, eye color body shape, and height, and many such more. It will depend on the genotype but also influenced by the factors existing in the surrounding. This article will enhance the concept of the difference between Phenotype and Genotype.
Table of content
Read Also: How To Calculate Time Physics
Advances Ambiguities And Open Questions
The experiments of Johannsen and Mendel can be seen as having achieved the goals given above .
Figure 2: Mendel-Johannsen method:Inbreeding, controlled crosses, and control of experimental conditionsallows unambiguous use of phenotypes to distinguish genotypes.
At the same time, Johannsen introduced many ambiguities andquestions about the import of his new terms. At first sight, the senseof classes is predominant. The phenotype, consisting of organismsdistinguishable by direct inspection or by finermethods of measuring or description , is used toidentify the genotype as a class of organisms that shares constituentsstable from generation to generation. Yet, no method is discussed todivide a natural varying population into phenotypes, let aloneidentify a genotype-as-class in such populations. It is in therestricted realm of inbred lines that identifying genotypes fromphenotypes is possible, albeit not reliably if a phenotype includes amix of inbred lines. Notice, however, if an inbred line is bred true, it is a genotype-as-class.There is no need to divide up the lines into phenotypes in order toidentify genotypes, and it matters not that the traits of individualsin an inbred line vary with the conditions in which the individualsare raised. Indeed, the norm of reaction of the inbred line is one wayto think of the genotype as an abstracted type.
2.2.1 Continuous Variation
2.2.2 Particulate Factors
2.2.3 Species-Shared Organization
Johannsen also noted that
What Counts Is Underneath Or Inside The Observable Surface
The genotype-phenotype distinction can also signify that thesurfacephenotypeis mere appearance what is underneathor inside that surfacethe genotypeis what counts. Asmall irony, given that the phenotype originated in relation toinferring genotypes , is that, to the extent that molecular biology has madeDNA sequences observable, especially at sites in which the sequencesvary from one group to another, each genotype becomes another phenotype . During the development of an organism, each of thesegenotypes-as-phenotypes at time 0 interacts with the rest of thephenotype and environmental factors to produce the phenotype at time1, 2, and so on. It may well be the case that germ cells arise at somepoint in the life course that are buffered from most of theseinteractions. However, with respect to conceptualizing developmentfrom time 0 till death, nothing logically makes the genotype not alsoa phenotype.
In any case, the view is widespread that what counts is underneath orinside. It is evident in the definition of evolution as change in genefrequencies and the idea that development of traits will eventually beunderstood in terms of a composite of the influences of DNA variantson the organism. It can also be seen in many other features ofdiscourses around heredity, such as the following:
Read Also: Easy Way To Learn Algebra Online Free
In Conclusion A Hint Of Epistemology
In its simplest expression, the epistemological concept of emergence states that the whole is more than the sum of its parts and more is different : a given organizational level may display properties that do not exist at lower levels. Biological systems, with their highly hierarchical organization, provide a myriad of examples. For instance, enzyme catalysis, rhythms, phyllotaxis spirals, the sense of smell, memory, consciousness, social organization, etc., are properties that emerge at a given level of organizationcellular, organismal or populationaland that do not make sense at other levels. The concept of phenotype is tightly linked to that of emergence. The phenotypic traits measured or calculated at a given organizational level characterize the properties that emerge at this level. Whether or not higher-level properties are reducible to lower level properties is a long-standing debate in the philosophy of sciences . Without taking sides, it is clear that a purely reductionist approachalbeit essential in biologyis not sufficient when dealing with properties at integrated levels, and systemic methods have to be used. This is the ongoing and daunting challenge we have to face to understand the genotypic bases of variation, a central question in biology.
Taxonomically Robust Gp Relationships
A mutation is expected to produce a somewhat reproducible phenotypic variation within a population. Such reproducibility in phenotypic outcome is required to allow genetic evolution and adaptation by natural selection . Indeed, a newly formed allele that would generate yet another phenotype each time it ends up in a different organism would not be subjected to natural selection. Reasoning in terms of variation, rather than considering alleles as isolated entities, makes it clear that competition occurs between alleles that span the same genetic locus. Natural selection acts directly on the allelic variation that is consistently associated with a given phenotypic variation, which is the GP relationship itself. The GP relationship is thus a basic unit of evolutionary change, on which natural selection acts.
Some of the most striking teachings of modern biology include the discovery that living beings share the same genetic material , the same genetic code , and the same basic cellular machinery. It is thus far from paradoxical that individual differences are built upon similarities, and the finding that certain GP relationships persists over long evolutionary times completes the picture.
Don’t Miss: Unit 1 Geometry Basics Homework 4
Several Current Representations Of The Connection Between Genotype And Phenotype Implicitly Dismiss The Differential View
We argued above that the differential view should always be kept in mind when thinking about the connection between genotypes and phenotypes. GWAS, which represent the most popular method to detect genomic loci that are associated with complex traits in populations, are based on the analysis of differences . Nevertheless, in current research the differential view is sometimes implicitly dismissed. When multiple factors are observed to influence phenotypic traits , the differential view is considered as too simplistic and researchers often prefer to focus back on phenotypes of single individuals, without explicitly relating them to a phenotypic reference.
FIGURE 2. Three current graphical representations of GP maps. The early version of the GP map proposed by Lewontin . A GP map where each point represents a single individual . The relationships between traits and genes, as depicted by Wagner . See text for details.
In another common graphical representation , a point in the G space and its corresponding point in the P space correspond to the genotype and the phenotype of a single individual . Under such a representation, the abstract object that we defined above as the GP relationship would correspond to a move in genotype space associated with a move in phenotype space . In a third representation put forward by Wagner , individual genes are connected to individual traits.
Shared Nature Of The Germ Cells Mechanics Of Development Material Basis For Genes
Some Mendelian researchers extended the investigation of the materialbasis for genes to their role in developmental processes. For example,the eyes of fruit flies, normally red, are sometimes white.Geneticists identified the location on the chromosomes thatcorresponds to the white-eye mutation and laterinvestigated the pigment-formation metabolic pathway and the enzymes involved as fruitfly eyes develop the normal or mutant color . Research since World War II that came to be known asmolecular genetics or molecular biology went on toidentify DNA as the chemical basis of genes and the mechanisms of DNAreplication, mutation, transcription to RNA, and translation topolypeptides . Researchers probed the feedbacknetworks that regulate these mechanisms, first in viruses andbacteria, then in complex, multicellular organisms mapped andmodified the specific DNA sequence of organisms compared sequencesamong taxonomic groups so as to assess the degree of geneticvariation in populations and to classify taxonomic groups intophylogenies traced where and when in development specific genes areactive and examined the role of DNA sequences not associated withgenes . Such research, which now occupiesthe center of biology, renders it plausible to many researchers andcommentators that development of traits will eventually be understoodin terms of a composite of the influences on the organism over time ofidentified DNA variants .
You May Like: First Day Of School Algebra 1 Activity
Classification Of Genetic Variants
Phenotypic variation in humans is a direct consequence of genetic variation, which acts in conjunction with environmental and behavioral factors to produce phenotypic diversity. Genetic variants are classified by two basic criteria: their genetic composition and their frequency in the population. In terms of composition, polymorphisms can be classified as sequence variants or structural variants. Sequence variants range from single nucleotide differences between individuals to 1 kilobase -sized insertions or deletions of a segment of DNA . Larger insertions and deletions, as well as duplications, inversions and translocations, are collectively called structural variants. These variants can range in size from 1 kb to those spanning more than 5 megabases of DNA .
Figure 1
Classification of genetic variants by composition. Schematic of sequence and structural variants compared to reference sequence. Sequence variation refers to single-nucleotide variants and small indels. Structural variation includes inversions, translocations and copy-number variants, which result in the presence of a segment of DNA in variable numbers compared to the reference sequence, as in duplications, deletions or insertions. Adapted from .
Figure 2
The Differential View Of Genotypephenotype Relationships
- 1CNRS, UMR 7592, Institut Jacques Monod, Université Paris Diderot, Paris, France
- 2Aix Marseille Université, CNRS, CEPERC UMR 7304, Aix en Provence, France
- 3Department of Molecular Cell Biology, University of California, Berkeley, CA, USA
An integrative view of diversity and singularity in the living world requires a better understanding of the intricate link between genotypes and phenotypes. Here we re-emphasize the old standpoint that the genotypephenotype relationship is best viewed as a connection between two differences, one at the genetic level and one at the phenotypic level. As of today, predominant thinking in biology research is that multiple genes interact with multiple environmental variables to produce the phenotype. Often, the problem of linking genotypes and phenotypes is framed in terms of genotype and phenotype maps, and such graphical representations implicitly bring us away from the differential view of GP relationships. Here we show that the differential view of GP relationships is a useful explanatory framework in the context of pervasive pleiotropy, epistasis, and environmental effects. In such cases, it is relevant to view GP relationships as differences embedded into differences. Thinking in terms of differences clarifies the comparison between environmental and genetic effects on phenotypes and helps to further understand the connection between genotypes and phenotypes.
Don’t Miss: What Does Expanded Form Mean In Math
Building A Phenomics Discovery Environment
How do we develop an environment in which researchers can readily make discoveries concerning the intimate connections among phenotypes, environment, and genetics? Three requirements must be met for this vision to become a reality across large-scale data. First, phenotype descriptions must be rendered in a computable format, which usually involves the use of appropriate ontology terms to represent the phenotypic descriptions found in narrative text or data sources. Each bit of text is thereby imbued with properties and relationships to other terms . Second, these semantically represented phenotype data, which integrate the phenotypes across species and also with their genetic and environmental contexts, must be stored in a way that is broadly accessible on the Internet in a nonproprietary format, e.g., in a Resource Description Framework . The third requirement is to grow a set of algorithms that enable users to analyze the data. That is, these algorithms combine the logical connections inherent in the ontologies with statistical analyses to, for example, identify similar phenotypes and their correlations with specific genetic or environmental factors.
Phenotypic Characteristics And Identical Twins
Another classical example of the environment’s influence on phenotype is in identical twins. Monozygotic twins have the same DNA sequences, hence the same genotype. They are not, however, phenotypically identical. They have phenotypic differences, in looks, behavior, voice, and more, that are observable.
Scientists have often studied identical twins to observe the environment’s impact on genes. Their identical genomes make them excellent candidates to help us decipher what else is involved in determining phenotype.
Two typical twin studies compare the following groups:
- Monozygotic vs dizygotic twins
- Monozygotic twins raised together vs. monozygotic twins raised apart.
Monozygotic twins come from the same original egg and sperm cells, which later on in the development process split to form two clumps of cells which eventually lead to two fetuses.
Dizygotic twins are from two different eggs and are essentially two siblings born in the same pregnancy. Thus, they are referred to as fraternal twins. They typically share about 50% of the same genes, while monozygotic twins share 100%.
When comparing monozygotic twins to dizygotic twins, scientists are attempting to discover phenotypic factors that are more heavily influenced by genetics. If all sets of twins were raised together, then any trait shared more heavily by monozygotic twins is a trait that has higher genetic control over phenotype.
Don’t Miss: Springboard Algebra 1 Unit 1 Answers
The Phenotypic Level Matters For The Genotypephenotype Relationship
As previously mentioned, traits can be measured and/or calculated at any level of phenotypic organization. Because trait variation is polygenic in the vast majority of cases, even for molecular phenotypes, the concepts and approaches of quantitative genetics can be applied, regardless of the level considered: quantitative trait loci are mapped and their effects measured, genetic effects such as dominance or epistasis are quantified, heritability is measured, etc. However, there are clear-cut differences between phenotypic levels regarding key genetic and evolutionary features:
Inbreeding depression is higher for life-history traits than for morphological traits. In a survey of 54 animal species, DeRose and Roff compiled inbreeding depression values for 35 life-history traits and 10 morphological traits such as adult body size, bristle number, etc. They showed that at \ , life-history traits experienced a mean reduction of \ in trait value, whereas morphological traits showed a mean reduction of only \. According to the authors, the most likely explanation is that positive dominance is on average lower for morphological traits than for life-history traits.