Crystallization Of The Sample Is Still An Major Hurdle
The major obstacle to the structural analysis of molecular assemblies has been and will continue to be in preparing suitable crystals. The growth of two-dimensional crystals has become a key aspect of electron microscopic structure determination, and new general approaches are urgently needed. Thus, the lipid-layer crystallization technique , if successfully developed, would play a critical role. The difficulties encountered in three-dimensional crystallization, as needed for highresolution x-ray analysis, will depend on the type of assembly in question. With membrane complexes, the crystals must be grown from precise mixtures of detergents, amphiphiles , protein, and lipid the process of crystallization has an additional dimension compared with that of soluble proteins. A major difficulty at present, therefore, is in obtaining sufficient commitments of financing and time to support such crystallization efforts. The first such crystallizations were carried out only after many years of trials in Europe, where the support of science can maintain a constant effort in a high-risk, long-term endeavor. Because risks have been demonstrably reduced, considerable weight must be given to early successes in growing crystals of sufficient quality for high-resolution analyses.
Structural Motifs Are Repeatedly Used To Carry Out Similar Functions
A structural motif composed of three extended strands and two helical coils of protein, named the nucleotide-binding domain, is found as part of many enzymes that bind nucleotides, such as adenosine triphosphate . Recently, from the nucleotide sequence of the gene encoding a protein associated with bladder cancer in humans, the nucleotide-binding domain was correctly predicted to be part of this protein’s three-dimensional structure.
Another structural motif has been seen in some of the proteins that recognize specific sequences of DNA and consequently regulate genes by turning them off or on. In this instance, two helical coils of protein connected by a short bend form a module that can plug into the major groove of a DNA double helix. The atomic surface of this recognition-helix motif is different in different proteins, imparting to them the ability to recognize and bind tightly to different specific sequences of DNA. As a result, one protein turns on one specific gene, whereas another might turn off another gene. Understanding how this structure functions has allowed scientists to synthesize novel regulators of genetic information.
Four Classes Of Biological Macromolecules
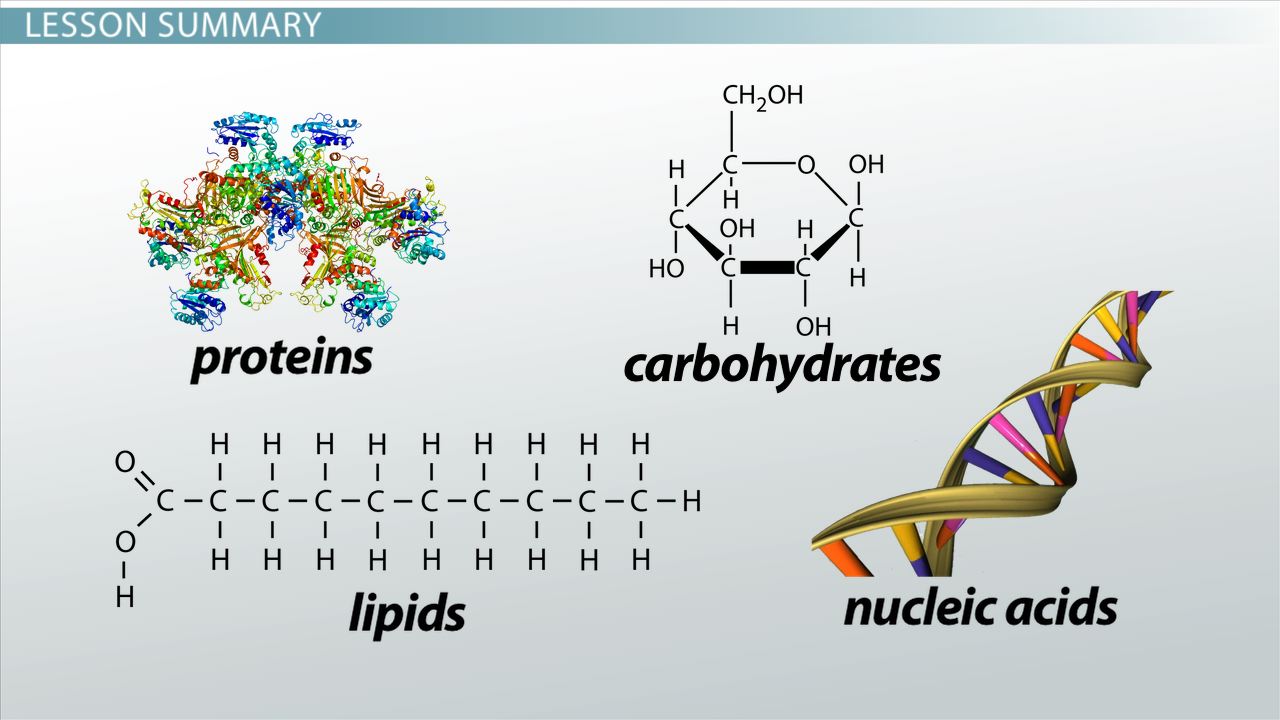
There are four major classes of biological macromolecules:
Each of these types of macromolecules performs a wide array of important functions within the cell a cell cannot perform its role within the body without many different types of these crucial molecules. In combination, these biological macromolecules make up the majority of a cells dry mass. All the molecules both inside and outside of cells are situated in a water-based environment, and all the reactions of biological systems are occurring in that same environment.
Interactive: Monomers and Polymers
Boundless vets and curates high-quality, openly licensed content from around the Internet. This particular resource used the following sources:
Boundless.
You May Like: Difference Between Inertial And Non Inertial Frame Of Reference With Examples
The Biological Building Blocks
The cell is the basic unit of life. All organisms are composed of one or more cells. As will be discussed later, humans are made up of many millions of cells. In order to understand what goes wrong in cancer, it is important to understand how normal cells work. The first step is to discuss the structure and basic functions of cells.
First we will introduce the common building blocks of cells. All cells, regardless of their function or location in the body, share common features and processes. Amazingly, cells are comprised almost entirely of just four basic types of molecules. Shown above is a cell surrounded by examples of these building block molecules.
Since they are present in living things these building blocks are called biomolecules. The next sections describe the structures and functions of each of these basic building blocks. Further information on the topics on this page can also be found in most introductory Biology textbooks, we recommend Campbell Biology, 11th edition.1
Dna And Rna Structure
DNA structure is dominated by the well-known double helix formed by Watson-Crick base-pairing of C with G and A with T. This is known as B-form DNA, and is overwhelmingly the most favorable and common state of DNA its highly specific and stable base-pairing is the basis of reliable genetic information storage. DNA can sometimes occur as single strands or as A-form or Z-form helices, and occasionally in more complex 3D structures such as the crossover at Holliday junctions during DNA replication.
RNA, in contrast, forms large and complex 3D tertiary structures reminiscent of proteins, as well as the loose single strands with locally folded regions that constitute messenger RNA molecules. Those RNA structures contain many stretches of A-form double helix, connected into definite 3D arrangements by single-stranded loops, bulges, and junctions. Examples are tRNA, ribosomes, ribozymes, and riboswitches. These complex structures are facilitated by the fact that RNA backbone has less local flexibility than DNA but a large set of distinct conformations, apparently because of both positive and negative interactions of the extra OH on the ribose. Structured RNA molecules can do highly specific binding of other molecules and can themselves be recognized specifically in addition, they can perform enzymatic catalysis .
Don’t Miss: What Does Abiotic Mean In Biology
Sequence Comparisons Lead To Structural Functional And Evolutionary Insights
Much valuable comparative sequence information awaits us as the data accumulate and as analytic methods become more reliable and informative. Already, one can do much using the data bases to help interpret any DNA sequence plucked more or less at random from a genome. The patterns of sequence in the regions that code for the amino acid chains of proteins differ enough from the noncoding regions that the former can usually be identified. For example, we know about types of sequences that are required for efficient synthesis of proteins in many different types of organisms. We know about some general types of control elements for certain genes important in developmental pattern formation or in an organism’s response to environmental stress.
Many proteins with related functions have probably evolved from common ancestors. Thus receptorsproteins designed to sit at the Cell surface and detect the environmentmay represent one or more fundamental families of structures and sequences. For example, the sequence of the beta-adrenergic receptor, which binds the hormone adrenalin, and the sequence for rhodopsin, which detects light, are sufficiently similar that we can tell both were once related through a common progenitor. In the same way, proteases often resemble other proteases and structural proteins resemble other structural proteins.
Nuclear Magnetic Resonance And X
NMR and x-ray diffraction provide both overlapping and distinct information about molecules. Recently, the chain-folding of a small protein was determined from an analysis of interatomic distances provided by NMR. X-ray diffraction simultaneously verified the structure, confirming as a side benefit that the structure of a protein in solution as seen by NMR is the same as that in a crystal as seen by x-ray diffraction. We can now confidently predict that NMR will make it possible to determine a series of structures of small proteins in solution.
You May Like: Movement Geography Definition
Dna Is An Dynamic Molecule That Can Switch Between Different Structural States
When the double helix structure for DNA was first announced, it was an instant public success. It represented a neat solution to a number of chemical and biological problems, and it was easy to describe and to remember. The importance of pairing between bases on the two DNA strands and stacking of adjacent bases along each individual DNA strand is overwhelming in nucleic acid structures. In terms of relative importance to the overall structure, there are no counterparts in proteins. However, with time, the structure of DNA has been found to be much more complex than was originally thought, since there are a variety of different double helical structures. The diversity of such structures has dramatically altered our thinking about the DNA molecule.
The dynamic aspects of the equilibrium structures of DNA have become clear with direct experimental measurement of the swinging in and out of individual bases to and from the axis of the helix. Larger scale motions on a much longer time scale are revealed by pulsed field gel electrophoresis, which separates molecules of enormous molecular weight.
Assembling And Disassembling Polymers
While there is variation among the types of biological polymers found in different organisms, the chemical mechanisms for assembling and disassembling them are largely the same across organisms.
Monomers are generally linked together through a process called dehydration synthesis, while polymers are disassembled through a process called hydrolysis. Both of these chemical reactions involve water.
In dehydration synthesis, bonds are formed linking monomers together while losing water molecules. In hydrolysis, the water interacts with a polymer causing bonds that link monomers to each other to be broken.
Also Check: How To Find Ksp Chemistry
The Protein Data Bank Is An Rich Resource For Predicting Structure
In the ad hoc approaches, the protein data bank is searched for patterns and statistical correlations. For example, probabilities based on the occurrence of each amino acid in various types of secondary structure differ and can, in turn, be used predictively to estimate probable regions of alpha helix, beta strand, and beta turn structures in any sequence. In parallel efforts, combinatorial algorithms aimed at packing assigned secondary structures into supersecondary and larger tertiary units have been developed. Most recently, combinations of such secondary and tertiary prediction schemes that show great promise in providing probable domain structures have been worked out. Whether the resulting models are close enough to converge to the native structure through molecular dynamic or energy minimization procedures is not yet known. Although all ad hoc approaches are implicity based on the underlying chemistry through the use of known structures, only a few explicity refer to these properties in the algorithm itself.
Genetically Engineered Proteins Reveal Much About How Proteins Function
The use of site-directed protein modification offers great promise for answering some of the fundamental questions in contemporary biology. For example cell-surface receptors must migrate throughout the cell from one organelle to another, moving from the endoplasmic reticulum to the Golgi complex to the plasma membrane . Once inside a coated pit, these proteins are taken inside the cell in a coated vesicle and then recycled back to the cell surface in a recycling vesicle. All of these movements seem to be dictated by signals contained within the structure of the protein itself. What are these targeting signals? Are they simply short, continuous stretches of amino acids or are they determined by the three-dimensional structure of the protein? Are protein modifications, such as phosphorylation or fatty acylation of the protein, required for any of these targeting signals?
The use of chimeric proteins has made it possible to define the functions of linear sequences responsible for protein translocation into the endoplasmic reticulum, mitochondria, and nucleus. However, signals that are defined by noncontinuous amino acid sequences are more difficult to define functionally with chimeric proteins. Incorrect protein folding becomes a major obstacle when the function of an internal sequence or domain is examined by this approach.
Read Also: Does Mj Have Any Biological Kids
Who Discovered The Structure Of Dna
The discovery of DNAs double-helix structure is credited to the researchers James Watson and Francis Crick, who, with fellow researcher Maurice Wilkins, received a Nobel Prize in 1962 for their work. Many believe that Rosalind Franklin should also be given credit, since she made the revolutionary photo of DNAs double-helix structure, which was used as evidence without her permission.
New Technology Has Improved Our Ability To Determine Complex Structures
These successes have stemmed in the main from technological developments and breakthroughs. In x-ray crystallography, particularly of viruses, the progress can be traced to four advances: the introduction of methods to acquire and process high-resolution data from very large repeating units the development of methods that take advantage of the symmetry of many molecular assemblies in solving the structure the tremendous advances in the speed, size, and affordability of computers and the development of computer graphics that efficiently communicate the information obtained.
An additional major breakthrough was learning to crystallize membrane protein assemblies in the form of protein-detergent micelles. Crystallization has been achieved now for several such membrane proteins, including the photosynthetic reaction center, E. coli matrix porin, and bacteriorhodopsin. Also of particular significance for membrane structure has been the progress made in electron microscopy. Preparative methods that preserve specimens have been developed as have mathematical analyses of the image that can provide three-dimensional details. The advent of cryo-electron microscopy, in which wet specimens are prepared by being frozen so rapidly that the water remains amorphous rather than crystalline, has made it possible to see functional molecules in action for the first time.
You May Like: Who Are The Biological Parents Of Prince Paris And Blanket
Biological Polymers: Proteins Carbohydrates Lipids
- B.A., Biology, Emory University
- A.S., Nursing, Chattahoochee Technical College
Biological polymers are large molecules composed of many similar smaller molecules linked together in a chain-like fashion. The individual smaller molecules are called monomers. When small organic molecules are joined together, they can form giant molecules or polymers. These giant molecules are also called macromolecules. Natural polymers are used to build tissue and other components in living organisms.
Generally speaking, all macromolecules are produced from a small set of about 50 monomers. Different macromolecules vary because of the arrangement of these monomers. By varying the sequence, an incredibly large variety of macromolecules can be produced. While polymers are responsible for the molecular “uniqueness” of an organism, the common monomers are nearly universal.
The variation in the form of macromolecules is largely responsible for molecular diversity. Much of the variation that occurs both within an organism and among organisms can ultimately be traced to differences in macromolecules. Macromolecules can vary from cell to cell in the same organism, as well as from one species to the next.
What Biomolecules Are Found In The Cell Membrane
Cell membranes contain a variety of biological molecules, notably lipids and proteins. Material is incorporated into the membrane, or deleted from it, by a variety of mechanisms:
The cell membrane consists of three classes of amphipathic lipids: phospholipids, glycolipids, and sterols. The amount of each depends upon the type of cell, but in the majority of cases phospholipids are the most abundant. In RBC studies, 30% of the plasma membrane is lipid.
Plasma membranes also contain carbohydrates, predominantly glycoproteins, but with some glycolipids . For the most part, no glycosylation occurs on membranes within the cell rather generally glycosylation occurs on the extracellular surface of the plasma membrane. The glycocalyx is an important feature in all cells, especially epithelia with microvilli. Recent data suggest the glycocalyx participates in cell adhesion, lymphocyte homing, and many others. The penultimate sugar is galactose and the terminal sugar is sialic acid, as the sugar backbone is modified in the golgi apparatus. Sialic acid carries a negative charge, providing an external barrier to charged particles.
The cell membrane has large content of proteins, typically around 50% of membrane volume.These proteins are important for cell because they are responsible for various biological activities. Approximately a third of the genes in yeast code specifically for them, and this number is even higher in multicellular organisms.
Don’t Miss: What Is Figure Ground Perception Psychology
What Are The Four Organic Molecules Found In Living Things
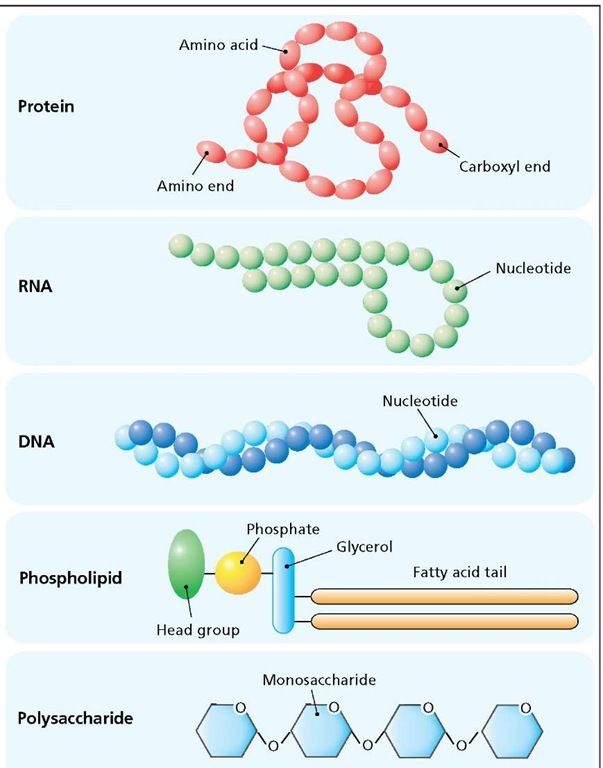
Living things are made of four types of molecules, known as macromolecules. These macromolecules are proteins, nucleic acids , lipids and carbohydrates. Each type of macromolecule is made of its own building blocks, which are intricately connected to form different shapes.
The special properties and shape of each kind of macromolecule are what make it especially suited for what it does. Proteins are machines that make and break other molecules. Nucleic acids carry genetic information that can be passed down to offspring. Lipids form barriers against water. Carbohydrates can be easily broken down for energy.
TL DR
There are four macromolecules that make up living organisms: proteins, nucleic acids, fats and carbohydrates.
Dehydration Synthesis Is How Monomers Become Polymers
In living cells, combining monomers into polymers occurs through a process called dehydration synthesis. The synthesis part of that term is straightforward: to synthesize means to put together, which is whats happening when monomers become polymers. Its called dehydration synthesis because a water molecule is removed from the monomers that are being combined together. Whos combining them? Well study this much more in a later module, but you should know for now that when monomers are combined into polymers, the agents doing the combining are biological catalysts called enzymes.
Heres a visual representation of a dehydration synthesis reaction.
An enzyme combines the two monomers. As it does, a water molecule is created as an OH is removed from the monomer on the left and an H is removed from the monomer on the right.
Heres another look at a dehydration synthesis reaction.
The two reacting molecules are amino acids, the monomers of proteins that we discussed above. Just as in the cartoon version of a dehydration synthesis above, note that on the right side of the first amino acid is an OH, and that on the left side of the second amino acid is an H . Both of these are colored blue to make it easy for you to find them. As these two amino acids are linked together, a water molecule is removed.
Also Check: Geometry Segment Addition Postulate Worksheet
Structure Is Determined By Several Factors
Covalent and non-covalent bonding govern the three dimensional structures of proteins and nucleic acids which impacts function. The amino acid sequences observed in nature are highly selected for biological function but do not necessarily adopt a unique folded structure. The structure of macromolecules is governed by foundational principles of chemistry such as: covalent bonds and polarity, bond rotations and vibrations, non-covalent interactions, the hydrophobic effect and dynamic aspects of molecular structure. The sequence of proteins and nucleic acids can be altered by alternative splicing, mutation or chemical modification. Sequences of macromolecules can evolve to create altered or new biological activities.
Associated learning goals