Value Of Tree Of Life
The tree of lifes value to the well-being of humans and existence is immense. The species cannot strive not to include a biodiverse planet. First, we need a complete biodiverse system in our environment to survive a strong life. For the sake of living organisms, biodiversity assumes a key role in all the subjects of life, that is, starting from food, medicines, and all other aspects of life for the survival of humans. Along with the aspects, it is also needed to understand the relationship between the living organisms that can be done only by understanding and relating biodiversity with the tree of life. Knowledge of this can also lead to various studies which may include ecology and its conservative measures.Knowledge of biodiversity and the tree of life could also lead to the mysterious natures discovery and its organisms along with the inventions of new and advanced medicine for the betterment of society. To know the relationships between them, we have to observe the habitat, temperature, and Change of factors according to the environmental conditions.Another major application of the tree of life is Phylogeny. Phylogenetic analysis also helps in discovering many diseases using the strains of viruses that were lost years ago. Phylogenetic DNA can be produced to represent these strains which help in the diagnosis of the diseases. For example, the flu occurs, using the phylogenetic strains we can be able to produce new flu vaccines using the strains.
Carl Woese And The Phylogenetic Tree
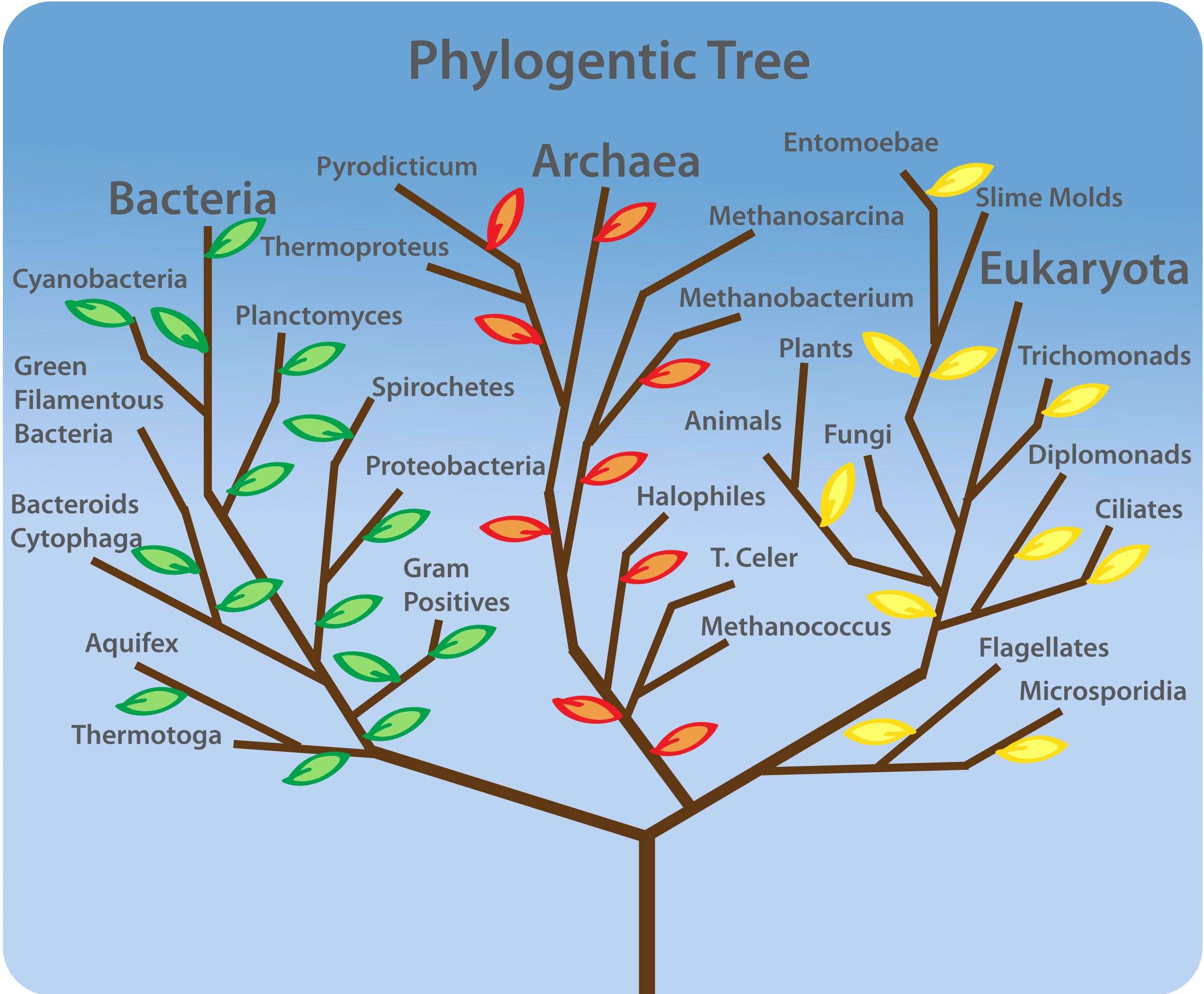
In the past, biologists grouped living organisms into five kingdoms: animals, plants, fungi, protists, and bacteria. The organizational scheme was based mainly on physical features, as opposed to physiology, biochemistry, or molecular biology, all of which are used by modern systematics. The pioneering work of American microbiologist Carl Woese in the early 1970s has shown, however, that life on Earth has evolved along three lineages, now called domainsBacteria, Archaea, and Eukarya. The first two are prokaryotic cells with microbes that lack membrane-enclosed nuclei and organelles. The third domain contains the eukaryotes and includes unicellular microorganisms together with the four original kingdoms . Woese defined Archaea as a new domain, and this resulted in a new taxonomic tree. Many organisms belonging to the Archaea domain live under extreme conditions and are called extremophiles. To construct his tree, Woese used genetic relationships rather than similarities based on morphology .
The Open Tree Of Life
So how can we keep track of whats out there and how its all related? Thats the problem of what we call big data. But strides are being made. A team of scientists recently published a digital database of life, called The Open Tree of Life, in the Proceedings of the National Academy of Sciences.
To create the database, they collected phylogenetic data from hundreds of published studies. Each study was of one small group of living things. Some of these studies did not come with digital databases, and when they did, they were of very different types. More than just organizing all the information into one useable database, scientists created this visual tree that shows the interrelationships of living things:
via CBSnews.com
This is the first real attempt to connect the dots and put it all together. Think of it as version 1.0, said principal investigator Karen Cranston of Duke University. Over the years, scientists will add to this, and the tree will grow.
Don’t Miss: Cyte Prefix Meaning
Trees Of Cells And Species
If one assumes, almost as an entailment of Schleiden and Schwanns cell theory, that all cells derive from previous cells, then it could be the case that there is a single tree-like pattern relating all cells, although one would need to look up to the level of species in lineages where sex became a reproductive necessity. Although incongruence of gene trees due to LGT means that no single one can be taken as topologically equivalent to such a tree of cells and species, the STOL might still be the best proxy for it. Patrick Forterre endorses this common view when he writes:
The universal tree should depict evolutionary relationships between domains defined according to the translation apparatus reflecting the history of cells and not according to the global genome composition that is influenced by LGT, virus integration and endosymbiosis, the history of which is incredibly complex .
Evolutionary Trees As A Representation Of Evolution
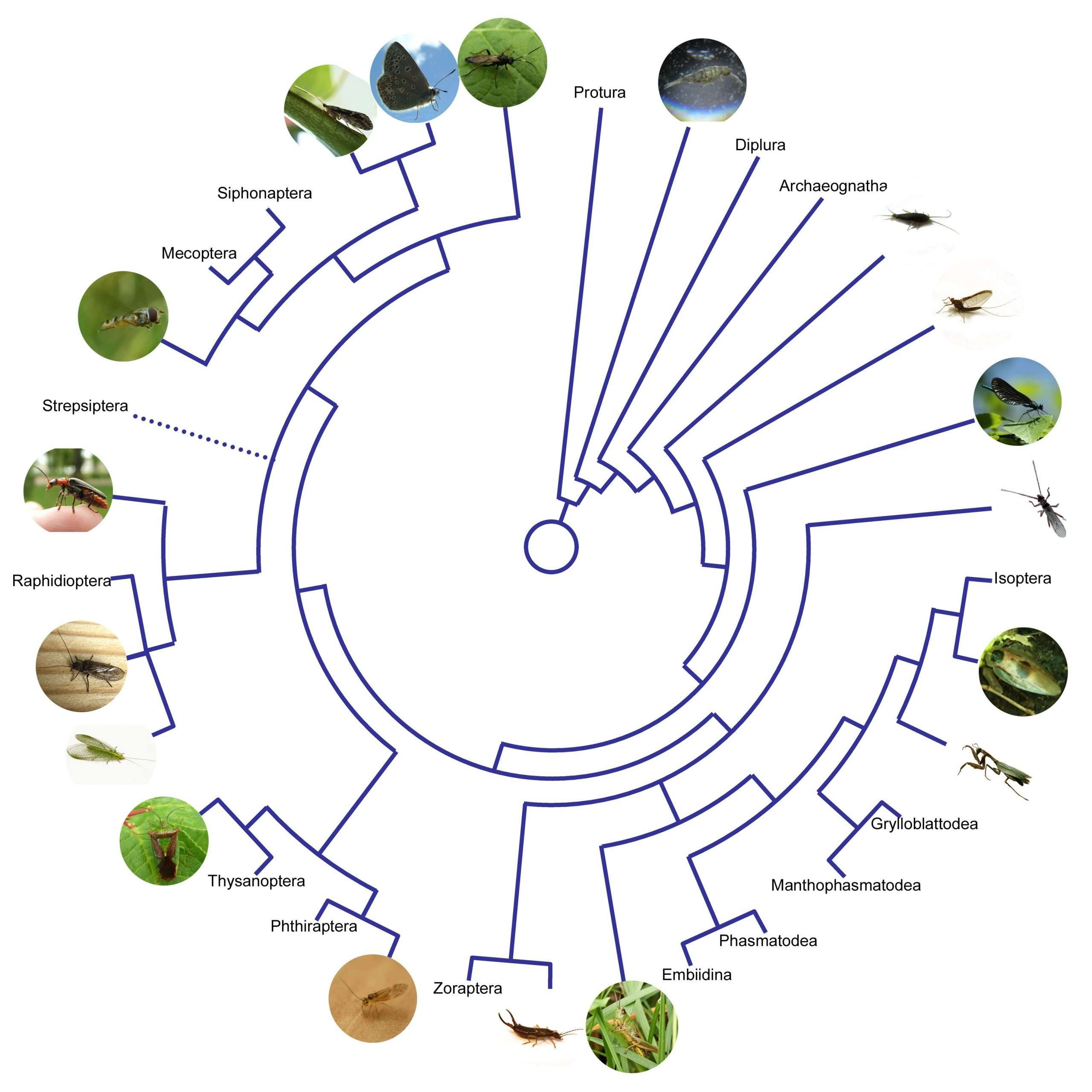
Evolutionary diagrams are diagrammatic depictions of different species or groups and their relatedness. They can exist in different forms , although treelike depictions are the most common . Crucial for evolutionary diagrams is that they present the evolutionary development of species and groups rather than the development or relatedness of individual organisms. In this paper, we will treat terms like evolutionary tree, phylogenetic tree, and phylogeny as interchangeable, although some experts in the field might use them with different connotations.
Evolutionary trees are a very common way to visualize patterns of macroevolutionary processes and therefore play a central role in teaching evolution . They serve as a tool to depict multiple relationships in one diagram, thus presenting processes and developments that are hard to describe in a simple way . In order to understand how evolution works, one has to understand how cladogenesis works, which is facilitated by knowing how speciation processes are represented in diagrams .
Fig. 1
Non-systematized tree-thinking skills
Non-hierarchical tree-reading framework
A hierarchical framework for tree-thinking
Halverson and Friedrichsen developed a hierarchical framework for tree-thinking skills. The hierarchy is based on the work of Kozma and Russell in the field of chemistry education and presents a skill set for the use of visual representations.
You May Like: What Causes Parallax Error And How Do You Avoid It
Hierarchical Relationships Between Groups Of Organisms
To the best of our current knowledge, all organisms that are alive today or that have lived on this planet in the past are part of one large, genetically connected group: Life on Earth. The three major groups of living things, Eubacteria, Archaea, and eukaryotes, are thus subgroups of the containing group Life on Earth. Each of these major subgroups of Life is itself divided into a multitude ofhierarchically nested subgroups. For example, eukaryotes is the containing group for a variety of groups including plants, animals, and fungi animals is the containing group for several groups including sponges, cnidarians, and Bilateria Bilateria is the containing groups for many groups like arthropods, molluscs, and nematodes, etc.
Life on Earth can be divided into a series of hierarchically nested subgroups, starting at the root of all life and ending at the tips in groups that cannot be further subdivided into distinct genetic lineages, e.g., Homo sapiens .
Links Between Tol Pages
In order to put the information about individual groups in a phylogenetic context, ToL branch pages feature a tree diagram . Tree diagrams provide an overview of the phylogenetic relationships among subgroups. For example, the tree diagram of the beetle page looks like this:
The tree diagram on the beetle page showing the relationships between the major beetle subgroups.
The basal branching point in the Coleoptera tree represents the ancestor of all beetles. This ancestor diversified over time into several descendent subgroups, which are represented as internal nodes and terminal taxa at the tips of the beetle tree. Beginning at the root on the left, the diagram shows that the ancestral beetle lineage gave rise, on the one hand, to members of the extinct Paleozoic group Protocoleoptera, and on the other to the ancestor of the remaining beetles. This ancestor in turn split into a species that gave rise to the extinct family Permocupedidae, and another that gave rise to the remainder of the beetles. Further splitting of ancestral species would lead us to the terminal taxa in the diagram, where we see Polyphaga and Myxophaga on adjacent branches. The ancestral species of these two groups split in two to give rise to one species that was the ancestor of the Polyphaga, and another species that was the ancestor of the Myxophaga. Even though it isn’t shown on this diagram, the splits implied in this tree probably happened in the Permian period, before the time of the dinosaurs.
Read Also: How Much Physics Is On The New Mcat
What Does The Tree Of Life Show Biology
tree of lifelifebiologytree showsbiological
. Correspondingly, what was the function of the tree of life?
The tree of life or universal tree of life is a metaphor, model and research tool used to explore the evolution of life and describe the relationships between organisms, both living and extinct, as described in a famous passage in Charles Darwin’s On the Origin of Species .
Beside above, what is the tree of life and how does it differ from a family tree? All of these are of the same species. A tree of life, however, goes back to our nth ancestors to connect us to other species. In other words, family trees are on a generational level, trees of lives are on special level.
Also to know, what are the three major branches in the tree of life?
This helps us decide which organisms should be together on the same “branches” of the tree of life. For example, we now know that fungi are more closely related to animals than they are to plants. Using these tools, we now think the three main branches of life are Archaea, Eubacteria, and Eukaryotes.
What is the story of the tree of life?
The impressionistic story of a Texas family in the 1950s. The film follows the life journey of the eldest son, Jack, through the innocence of childhood to his disillusioned adult years as he tries to reconcile a complicated relationship with his father .
Nearly Universal Trees And The Statistical Tree Of Life
A decade and a half of prokaryotic tree-making has not produced general agreement on just how much LGT there is, other than much more than we expected, although there have been many serious attempts . Indeed, there is little agreement on how to quantify LGT as a process and its impact on genomes over all of evolutionary time. The most reasonable way, we think, is to ask of any contemporary genome how many of its genes have been strictly vertically inherited along a lineage of replicating genomes, tracing all the way back to that of the last universal common ancestor . There are only about 100 genes that are found in all or nearly all prokaryotic genomes and thus have a strong claim to having been present continuously in all lineages . These are largely involved with the ribosome and its functions , and are considered to be relatively immune to LGT, according to the complexity hypothesis .
Puigbo et al. have for some time been conducting large-scale gene family tree comparisons across bacteria and archaea, the Forest of Life project involving thousands of such families. Even among their 102 Nearly Universal Trees , made from genes found in all or nearly all bacteria and archaea, there are very few with identical histories. However, NUT topologies are far more congruent than expected by chance, so they appear to reflect a significant central trend, an attractor in the tree space that could be equated with the STOL .
Recommended Reading: Is Paris Jackson Michael Jackson’s Biological Daughter
Introduction: Why Are Evolutionary Trees Important
Graphical representations play a major role in modern learning environments. Various types of graphs, diagrams, charts, and schemata can be found in all areas of scientific research and various forms of learning materials . This applies in particular to biology, where students are regularly confronted with a large variety of different representational styles in its many subdomains .
Teaching biological fields like microbiology, genetics, and evolution involves teaching abstract processes that may not be observed by students. Furthermore, the concept of evolution is not intuitive . Evolution is the conjunctive core principle of biology and in order to develop a deeper understanding of any biological discipline, one needs to grasp the concepts of evolution . Its understanding requires knowledge of various different and seemingly unrelated topics like genetics, ecology, and morphology , and opportunities for practical work are rare . This leads to evolution being seen as one of the most challenging topics to teach in introductory science courses . The difficulty in teaching evolution is compounded by numerous and very persistent misconceptions prevalent among learners of all ages and education levels .
Reviewer : John M Logsdon Jr
The prokaryotic tree of life is dead!
The message rings clear in this extraordinary paper from an ensemble group of biologists and philosophers of science. In some ways, I am convinced–and others should be, too. That, I suspect, is the main objective of this paper: to provide the reader with an overwhelming “disproof” of the standard view that prokaryotic evolutionary history occurred as lineage-splitting events and can be depicted by a single bifurcating tree. By interweaving philosophical, technical and empirical arguments, a solid case can be made for inapplicability of traditional tree-thinking and tree-making to prokaryotes. But I also suspect that the larger goal is to simply challenge the readers’ deep-seated sensibilities that such trees must necessarily be at the heart of how we view evolutionary relationships of all organisms.
In sum, this thought-provoking paper may help to pave a clearer intellectual path for stubborn tree-monists like myself. Although the suggestion of possible successors to the traditional tree of life view is a positive step forward, I have a nagging feeling that in embracing pluralism we just might be missing the actual trees for the forest.
Long live the prokaryotic tree of life!
Answer to John Logsdon
Don’t Miss: Kuta Software Simplifying Radical Expressions Answers
Genes From The Expanded Marker Set Are Not Widely Distributed In Archaea
The 381-gene set was derived from a larger set of 400 genes used to estimate the phylogenetic placement of new lineages as part of the PhyloPhlAn method and applied to a taxonomic selection that included 669 Archaea and 9906 Bacteria . Perhaps reflecting the focus on Bacteria in the original application, the phylogenetic distribution of the 381 marker genes in the expanded set varies substantially , with many being poorly represented in Archaea. Specifically, 41% of the published gene trees contain less than 25% of the sampled archaea, with 14 and 68 of these trees including 0 or 10 archaeal homologues, respectively. Across all of the gene trees, archaeal homologues comprise 014.8% of the dataset . Manual inspection of subsampled versions of these gene trees suggested that 317/381 did not possess an unambiguous branch separating the archaeal and bacterial domains . These distributions suggest that many of these genes are not broadly present in both domains, and that some might be specific to Bacteria.
Process Pluralism And Its Implications For Taxonomy
All the above has important implications for the “species” notion as well. Rather than working under a single unified concept, microbiologists already accept many different pragmatic definitions of prokaryotic species. They have no species concept that would be relevant for all of life that would justify the reconstruction of a universal species tree. Doolittle and Zhaxybayeva showed that due to various genetic, population ecological, and evolutionary processes, not all prokaryotes belong to genomically and phenotypically cohesive clusters that biologists could be defined as “species” . In some instances, life-defining processes work together and generate groups of related organisms, sufficiently like one another to be called species. However, the evolution of such coherent clusters is not the general outcome in the prokaryotic world. Rather, various prokaryotic species taxa are defined in nature based on many different criteria, such as global genetic distance and the presence of some cohesion mechanism . Based on such criteria it is the case that there are multiple correct ways to classify the organic world, and a single organism may be classified in more than one manner depending on the aims of classification.
You May Like: Is Paris Jackson Michael’s Biological Kid
Biology And The Tree Of Life
Life 5 characteristics of living organisms Made up of cells Store and process heritable information They replicate Populations evolve Use energy
Cells Gamete cells, breast cancer cells Features Contain molecular information that encodes their physical attributes Have a boundary between them and the environment – allows for building up of chemicals Have the ability to harness materials and/or energy Concentrate reagents for biological reactions Make possible chemical gradients across the plasma membrane Link a phenotype to the same physical space that encodes it
Cell Theory A cell is a highly organized compartment bounded by a plasma membrane that contains concentrated chemicals in an aqueous solution All cells have a plasma membrane Within a multicellular individual, all cells are descended from a progenitor cell the zygote Zygote Derived from other cells Implications All cells come from preexisting cells, all individuals in a population of single-celled organisms are related by common ancestry In a multicellular organism, all of the cells present descend from preexisting cells and are also related by common ancestry We can represent many relationships among organisms with phylogenetic trees
Testing Cell Theory Maggots Observation: maggots appear in meat left out Null hypothesis: maggots arise spontaneously Appear in an open jar Do not appear in a gauze-covered jar
Finding Ancient Vertically Evolving Genes
The relationship between marker gene verticality, Archaea-Bacteria branch length, and functional category.
Vertically evolving phylogenetic markers have longer AB branches. The plot shows the relationship between a proxy for marker gene verticality, within-domain split score , and AB branch length for the 54 marker genes. Marker genes with higher split scores have shorter AB branch lengths . Split scores of ribosomal and non-ribosomal markers were statistically indistinguishable . Among vertically evolving marker genes, ribosomal genes do not have a longer AB branch length. The plot shows functional classification of markers against AB branch length using 54 vertically evolving markers. We did not obtain a significant difference between AB branch lengths for ribosomal and non-ribosomal genes .
Recommended Reading: What Effect Did Geography Have On The Way Greece Developed